Atlas of Variant Effects Alliance:
Precision medicine at nucleotide resolution
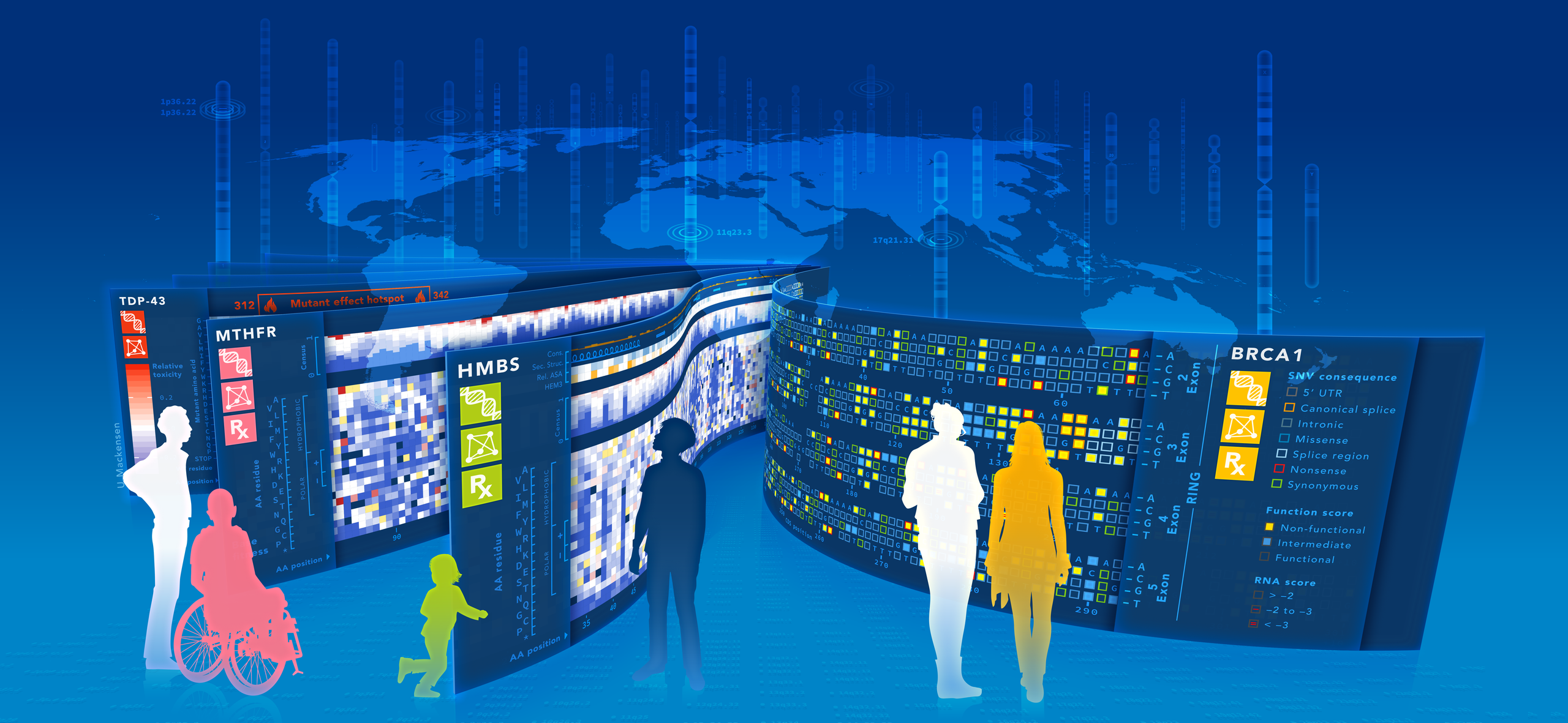

Art by Uta Mackensen (CC BY-ND) Image Description: Background: A world map and chromosome idiogram. Foreground: People moving amongst and inspecting larger than life Variant Effect Maps of clinically important genes BRCA1, HMBS, MTHFR and TDP-43.
The vision of the Alliance is to create comprehensive variant effect maps for important regions of human and human pathogen genomes that could ultimately assist in the diagnosis, prognosis and treatment of disease. The goal of our Alliance is to bring together data generators, curators and consumers, along with funders and other stakeholders, to set standards, share tools and develop strategy.
By describing the effects of variants in the genome, the atlas will accelerate and empower biological research, drug discovery and medical practice.
Graphic Credits: kjpargeter Freepik, Sayeh Gorjifard and Uta Mackensen
The Alliance welcomes individuals from academia, industry, government or other entities anywhere in the world
In this series, early-career scientists from around the globe share and discuss their research related to interpreting human genetic variation
Latest Event
Latest AVE Mention in the News
27 April 2026.
Nearly five years of consistent programming, a growing global community, and an enduring commitment to open science’, the Variant Effects Seminar Series (VESS), hosted online each month by the Atlas of Variant Affects (AVE) Alliance, will celebrate a milestone in June: its 100th presenter since it was launched in 2021.
Latest Seminars
7 July 2026.
Sarah Gurev is a FutureHouse AI-for-Science Independent Postdoctoral Fellow, co-advised by Sergey Ovchinnikov and Aaron Schmidt. She recently completed her PhD in Electrical Engineering and Computer Science at MIT in Debora Marks lab. She develops machine learning models to address unmet needs in virology and immunology, with an emphasis on pandemic preparedness and therapeutic development. Her research spans viral evolution and function, precision-engineered vaccine design, viral-host interactions, and immune receptor modeling.
2 June 2026.
I'm a postdoctoral researcher at the Dias and Frazer lab at the Centre for Genomic Regulation (CRG) in Barcelona, where I work on disentangling the biophysical effects of missense variants. My background is in physics and biological chemistry — I did my PhD at the University of Buenos Aires studying protein folding thermodynamics and evolution. I'm broadly interested in connecting evolutionary and physical principles to understand how mutations reshape protein behavior.
AVE in Action
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
9th Annual Mutational Scanning Symposium in Melbourne, Australia
Image credit: Ivan Martí
2025 Mutational Scanning Symposium (IBEC Barcelona Spain)
Image credit: Ivan Martí
2025 Mutational Scanning Symposium (IBEC Barcelona Spain)
Broad Mutational Scanning Symposium
Toronto Lab
Clare Turnbull and Shawn Fayer
MAVE workshop
MAVE workshop
Hasan and Marcin Marsh Lab
Cell Factories
Broad Mutational Scanning Symposium
Broad Mutational Scanning Symposium
Broad Mutational Scanning Symposium
Image credit: Greg Moss (Wellcome Sanger Institute)
Larissa Matsuyama, Fernanda Arriaga González and Rebeca Olvera León (SGE team, Welcome Sanger Institute)